Advanced R packaging
Building FAIR R packages for scientists
Program
- How to package (briefly)
- Quality of life setup
- Building FAIR R packages
- Packaging scientific code
How to package
All you need to know

Freely available at r-pkgs.org
Minimal package infrastructure
DESCRIPTION: Metadata (maintainers, dependencies, license)
man: Help pages*
NAMESPACE: Exported names*
R: R code
*usually automatically generated via devtools package
Tutorials
 e.g., carpentries and Utrecht University
e.g., carpentries and Utrecht University
Quality of life setup
All things that reduce mental load
Assuming the package is developed on GitHub
Development workflow: devtools

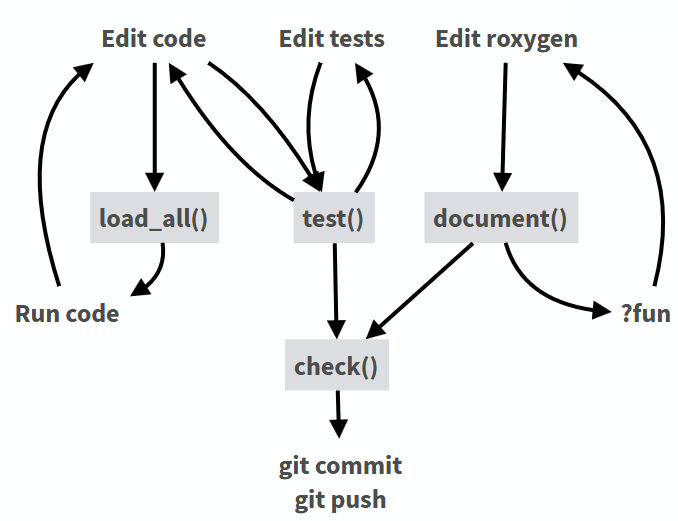
Spellchecking: spelling

- Adds spellchecker to
devtools::check() - Whitelist for technical terms
Unit Testing

Unit testing with testthat
Write tests:
Run tests:
Testing in general
- Many more options as part of
testthat:expect_true,expect_erroretc. - Wide range of tests available in other packages: Plots, server responses, “golden tests”, etc.
- Great to reduce cognitive load in complex packages - if something behaves unexpectedly you will know
- Not everything can be tested, especially in scientific software
- But: Writing tests influences design choice
Continuous Integration/Continuous Deployment (CI/CD)
Automatically test code across OSs and R versions
> usethis::use_github_action()
Which action do you want to add? (0 to exit)
(See <https://github.com/r-lib/actions/tree/v2/examples> for other options)
1: check-standard: Run `R CMD check` on Linux, macOS, and Windows
2: test-coverage: Compute test coverage and report to https://about.codecov.io
3: pr-commands: Add /document and /style commands for pull requests
Selection: Typically devtools::check() upon push to main
Building FAIR R packages
FAIR principles
FAIR principles for research software (Barker et al. 2022)
- Findable: Software & metadata has a PID
- Acessible: Can be retrieved via the PID
- Interoperable: Reads, writes and exchanges data in domain-relevant standards
- Reusable: Licensed, citable, contains provenance
Findable
Archived using a DOI
- Easiest setup: GH - Zenodo integration with automated archiving and DOI assignment upon release
- CRAN assigns DOIs, but persistence unclear
- GH is NOT an archive!
Accessible
Can be downloaded from a public repository
- CRAN is standard, your packages should always be there
- Domain specific alternatives (e.g., Bioconductor for bioinformatics)
- Conda forge?
- R-universe?
Interoperable
Reads, writes, and exchanges data in relevant standards
- Difficult with the R OO systems (S3)
- Document your classes and their expected fields
- Provide clear interfaces with other data formats
- Expose your internal functions! Scientific code is notoriously hacky
Reusable

- Provide a license (ask your funder or PI)
- Citation info in
CITATION.cffor underinst/CITATION - Generation via
cffr - Automatic transfer of metadata to archives
- Cite software in your publications
CI/CD meets FAIR
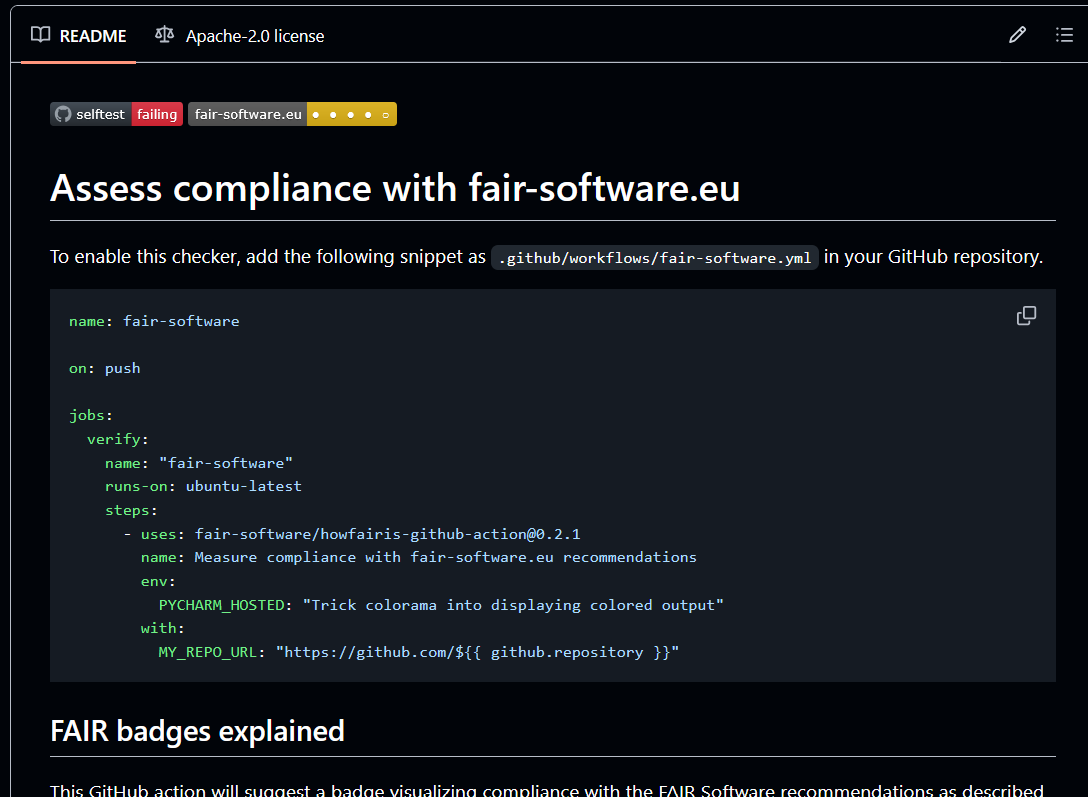
Why packaging (scientific) code?
Packages as scientific output

Software as part of Recognition & Rewards
Packages as scientific output: ECR edition
Packages can be part of a dissertation (talk to your supervisor!)
Develop skills for a career outside of academia
Packages as scientific infrastructure
FAIR data and software as key component of Open Science
Increasing focus on software sustainability
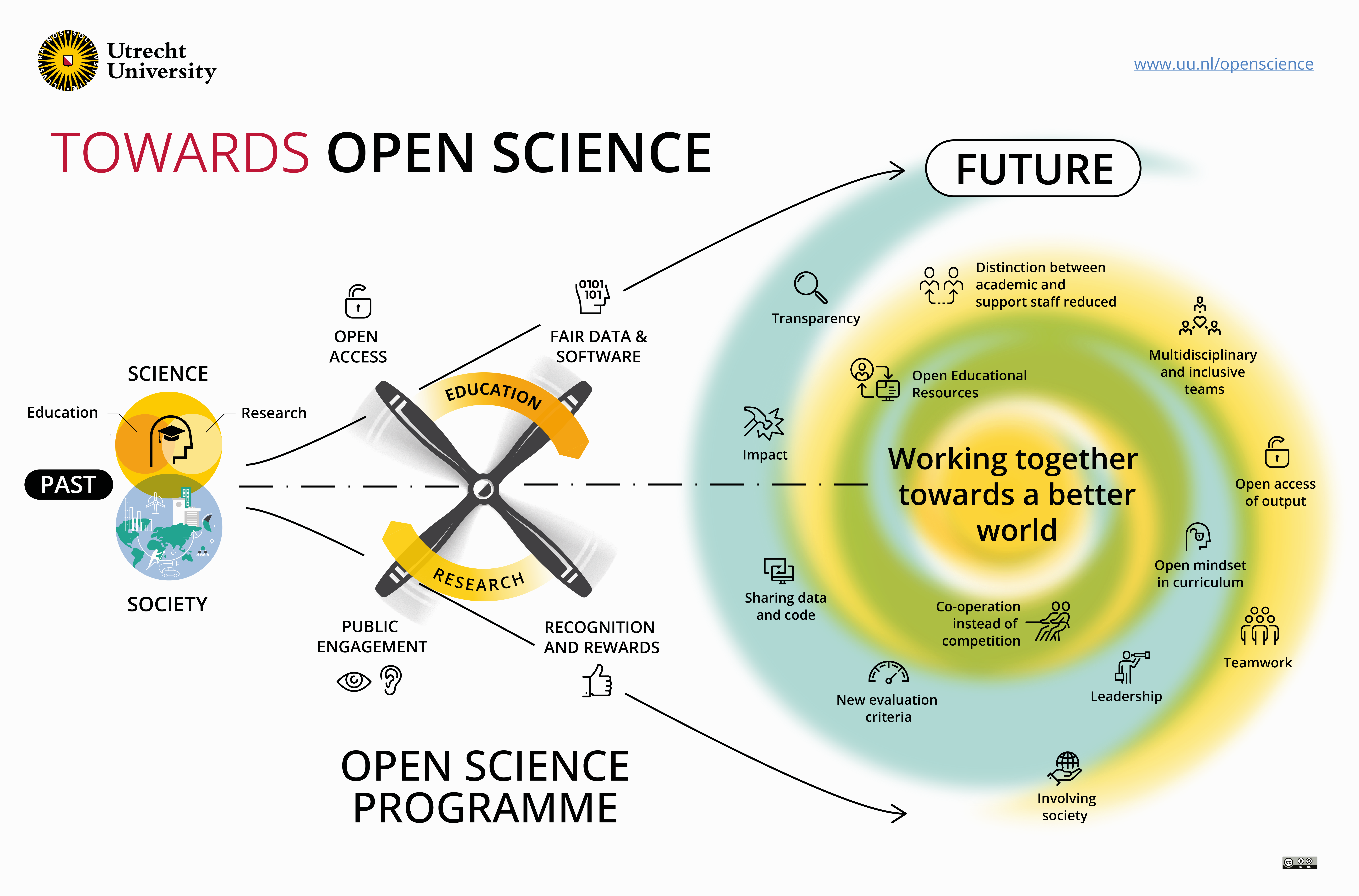
uu.nl/en/research/open-science
Community engagement
- Scientific communities are defined by the tools they use
- Sharing these tools helps build a community and promote methodological approaches (e.g., palaeoverse.org)
Minimum effort:
- Write a CONTRIBUTING.md
- Protocol for bug reports
- Think of governance after end of project
Website
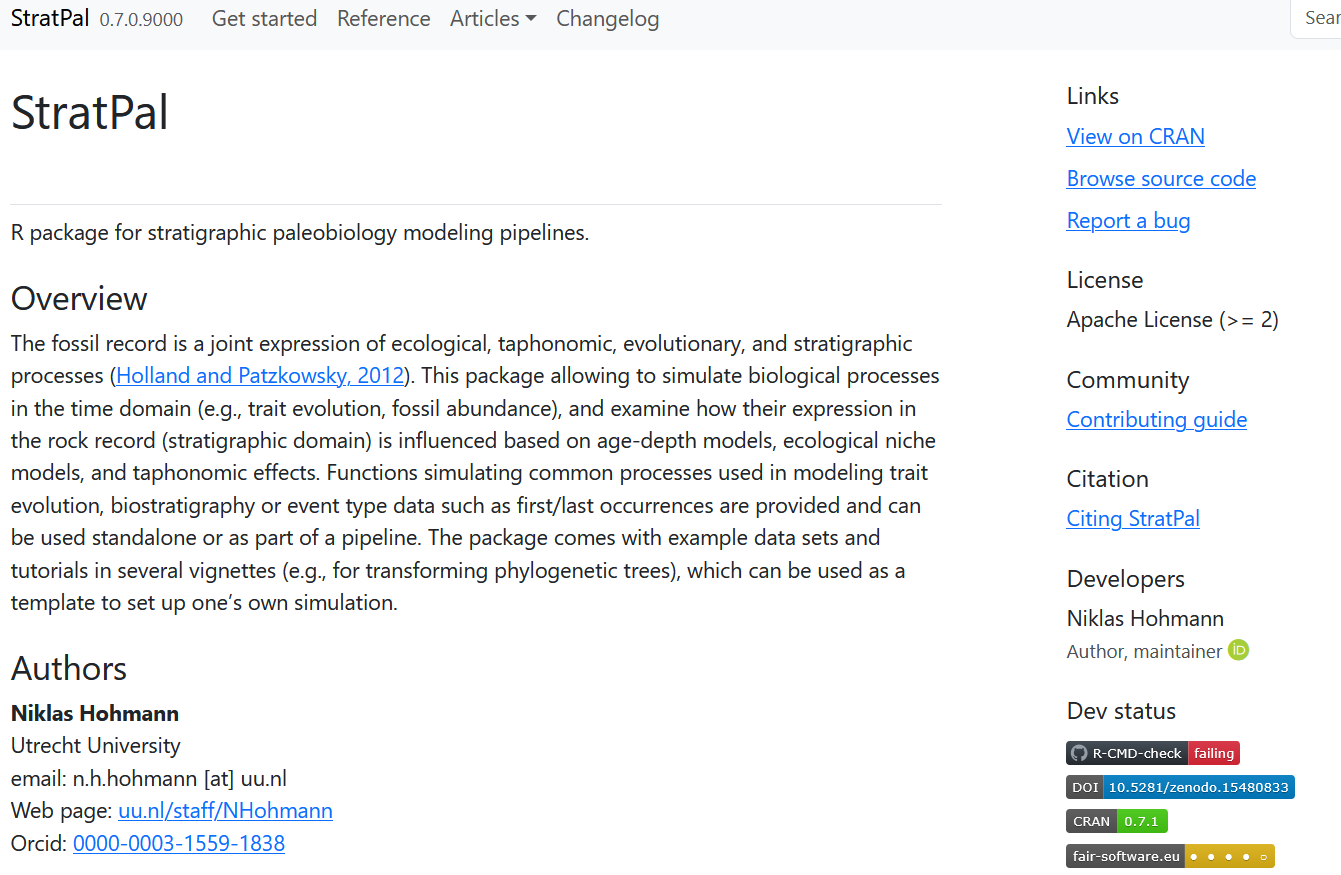
Combines documentation, vignettes, license, contribution guidelines, bug reporting and much more
Downsides of packaging
- Slower turnover
- Development and maintenance overhead
- Hard to get the community engaged & get contributions
- Software as scientific output not internationally recognized
Example: StratPal package
Links geosciences (stratigraphy) with biology (paleontology & phylogenetics)

StratPal publication

Teaching materials

Shiny App + open educational resources
Workshop materials
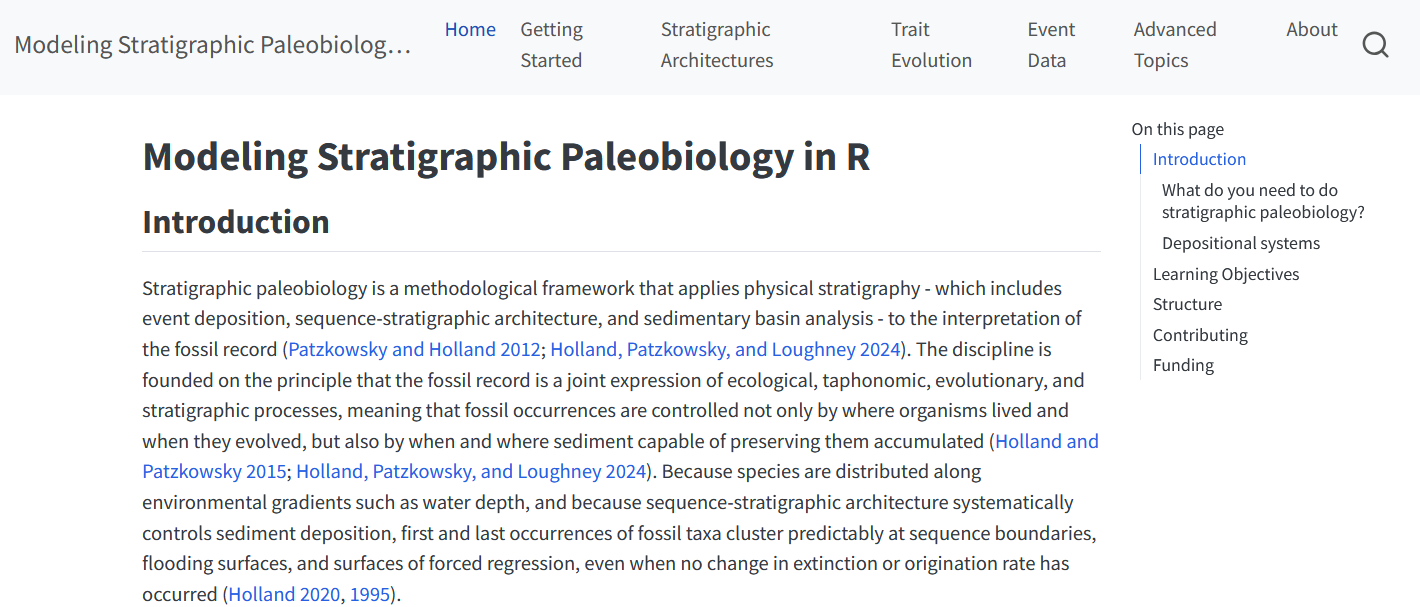
StratPal webpage

Summary
- Packaging is easy, with plenty of infrastructure available to reduce cognitive load
- Package your scientific code to share code with collaborators and across publications and engage your community
- Packages are standalone scientific outputs. Treat them with the same care as publications
- Be aware of the overhead that comes with developing and maintaining packages
Hands on
Got your own package?
- FAIRify - add license, make webpage, link to Zenodo, release on GH, … - whatever is most relevant to you
No package?
- Get started under github.com/UtrechtUniversity/tutorial-r-package

Niklas Hohmann